A team of scientists from the University of Chicago has managed to coax out the secrets of HIV budding, offering a new avenue of combating the frightful virus.
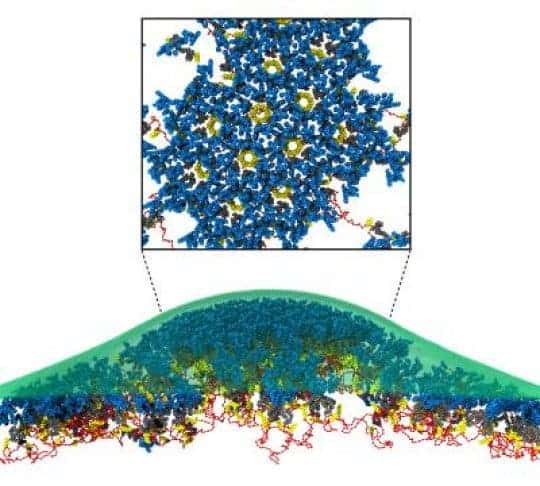
Image credits Voth et al., 2017, PNAS / University of Chicago.
HIV seems to have a particular regard for the Trojan horse approach to warfare. The process is known as budding and helps HIV infect cells while staying undetected by the body. After infecting a cell, the virus forces it to form a membrane capsule filled with more of the virus. When full, this capsule is released through “budding” and floats away. Upon contacting another cell, the capsule is allowed through its membrane then promptly falls apart, starting the process anew.
“[Budding is] the final step of seven steps in the HIV life cycle. During budding, immature (noninfectious) HIV pushes itself out of the host CD4 cell. (Noninfectious HIV can’t infect another CD4 cell.) Once outside the CD4 cell, the new HIV releases protease, an HIV enzyme. Protease acts to break up the long protein chains that form the noninfectious virus. The smaller HIV proteins combine to form mature, infectious HIV,” according to the U.S. Department of Health and Human Services.
However, one team of scientists at the University of Chicago is determined to take this weapon away from HIV’s arsenal. Through computer modeling, they were able to clarify previously unknown details about HIV budding. The findings could help us create a novel line of medicine to fight the virus, and offers a novel avenue of viral research in the future.
To gag a virus
It’s previously been determined that a key component of the budding process is a biochemical protein complex called Gag. However, the exact details of budding, as well as the exact structure of the Gag complex have remained largely unexplained. This prevented the development of medicine that could counteract this process.
“For a while now we have had an idea of what the final assembled structure looks like, but all the details in between remained largely unknown,” said Gregory Voth, the Haig P. Papazian Distinguished Service Professor of Chemistry and corresponding author on the paper.
Efforts to get a good image of what the protein complex looks like on a molecular level have sadly been unsuccessful so far. As such, Voth’s team turned to computer modeling to simulate Gag in action, and from there infer its properties and structure.
First, they built their model using known parts of the Gag complex. They then simulated the interactions of this model and the conditions within cells, fine-tuning it to match cellular infrastructure and synthesis capability. Progressive tweaking of the model allowed them to zero in on the most likely configuration of the protein and the process it supports.
The team then ran a battery of tests at the National Institutes of Health and the Howard Hughes Medical Institute Janelia Research Campus, overseen by co-author Jennifer Lippincott-Schwartz, to validate their findings. And it worked.
The findings offer hope that a novel range of medicine can be developed to counteract budding, severely limiting HIV’s ability to spread or remain undetected by the immune system. In concert with methods of boosting white cells’ ability to fight the virus, this could finally produce an effective, sure-fire cure against HIV.
Another exciting element of this study is that the team proved computer simulation can come in and fill the gaps in our understanding of viral mechanisms. In cases where direct observation of molecular processes just doesn’t work, this study offers a powerful precedent.
“The hope is that once you have an Achilles’ heel, you can make a drug to stop Gag accumulation and hopefully arrest the virus’s progression,” Voth says. “It really demonstrates the power of modern computing for simulating viruses.”
Next, the researchers plan to look at the Gag complex in the HIV capsules after budding, he adds.
The paper, “Immature HIV-1 lattice assembly dynamics are regulated by scaffolding from nucleic acid and the plasma membrane,” has been published in the journal Proceedings of the National Academy of Sciences.






