A group of researchers successfully sequenced the genome of the mold that produced penicillin, the world’s first true antibiotic, using samples frozen alive more than fifty years ago. The team compared Alexander Fleming’s original sample of penicillium mold to two strains of mold now used to produce the substance today.
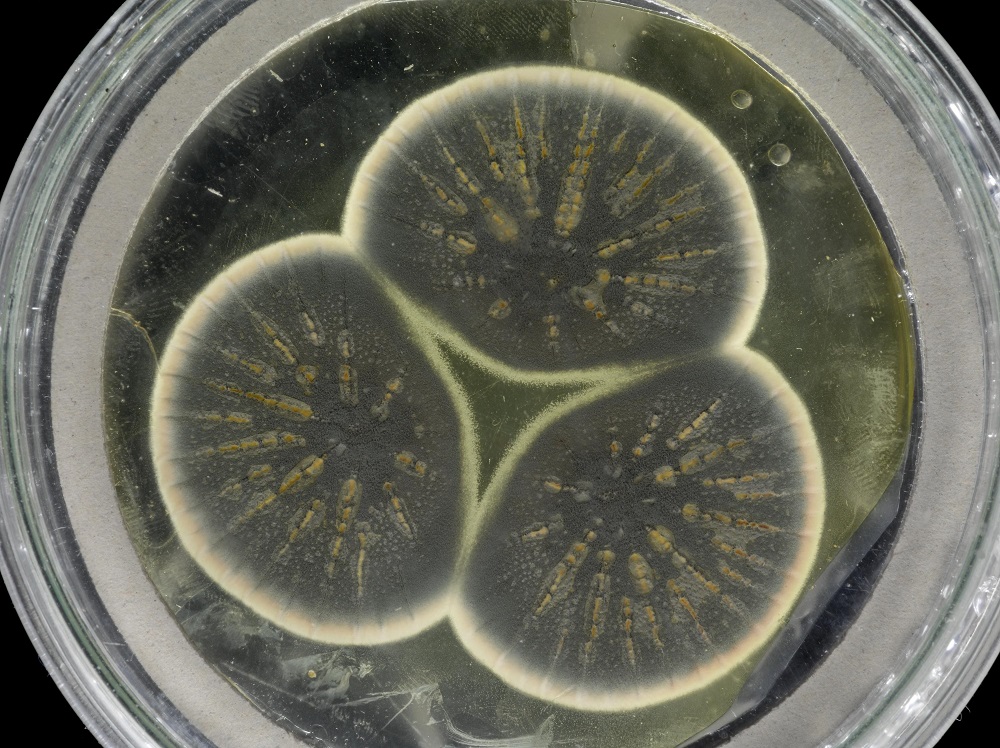
Back in 1928, biologist Alexander Fleming noticed Penicillium mold growing in a culture of Staphylococcus aureus he was studying. It appeared the experiment was wrecked but Fleming noticed that where the mold grew, the bacteria didn’t. He later identified the chemical compound that was fatal to the bacteria and called it penicillin in honor of the humble mold.
Fleming froze samples of the mold that produced his first isolated samples of pure penicillin. More than 50 years later, a group of researchers at Imperial College London and the University of Oxford decided to look them up. They compared the samples with the genomes of two modern strains of Penicillium mold, now used in the United States.
“We originally set out to use Alexander Fleming’s fungus for some different experiments, but we realized, to our surprise, that no one had sequenced the genome of the original Penicillium, despite its historical significance to the field,” said Timothy Barraclaugh, co-author, in a statement.
The researchers found a subtle difference between the two genomes, which might help us better combat antibacterial resistance. Most antibiotics are based on chemicals that fungi or bacteria produce to defend themselves. If you get a dose of penicillin, it was likely produced by mold cultures, which are descendants of samples taken from moldy cantaloupes.
Over the years, antibiotics manufacturers bred their cantaloupe mold cultures to produce more penicillin. This means the genomes of modern industrial Penicillum mold are probably very different from their cantaloupe-eating ancestors.
The team looked at two sets of genes in particular. The ones that coded for chemicals called enzymes and the ones that control how much of an enzyme to make and when. They found that modern strains had more copies of the genetic instructions for making those enzymes, which meant those cells would make more enzymes and thus more penicillin.
While nature favors the traits that make mold more likely to survive and pass on its genes, artificial selection by humans cares about penicillin production over everything else. But Fleming’s mold and the modern strains used different versions of the enzymes that make penicillin. This could be due to evolution in the lab or because the strains are from different continents and evolved different enzymes.
If that’s the case, those different enzymes might produce different versions of penicillin. Still, there’s not enough data now to say exactly how the different enzymes impact the final product. The difference could lead to more efficient penicillin production, more effective penicillin, or a way to work around at least some of the resistance certain bacteria strains have evolved to the drug, the researchers believe.
“Industrial production of penicillin concentrated on the amount produced, and the steps used to artificially improve production led to changes in numbers of genes,” Ayush Pathak, lead author, said in a statement. “But it is possible that industrial methods might have missed some changes for optimizing penicillin design, and we can learn from natural responses to the evolution of antibiotic resistance.”
The study was published in the journal Scientific Reports.






