It’s a game changer – scientists have discovered a new class of antibiotics which can kill an array of germs by blocking their capacity to build their cell walls, making it extremely difficult for bacteria to evolve resistance. It’s the first such discovery in the past three decades, and comes as a much needed breath of air in the fight against superbugs.
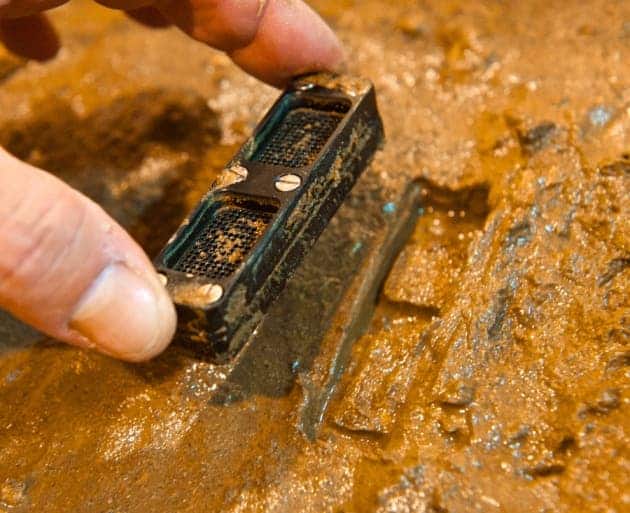

The antibiotic is called teixobactin and was uncovered by screening 10,000 bacterial strains from soil. Teixobactin has been tested on animals, but it’s still yet to be tested on humans. The results were described in Nature.
New antibiotics, new prospects
“Teixobactin kills exceptionally well. It has the ability to rapidly clear infections,” said research leader Kim Lewis, director of the Antimicrobial Discovery Center at Northeastern University in Boston, US.
The discovery of this antibiotic is exciting from many reasons – first of all, it’s been quite a while since a new class of antibiotics has been developed. Second of all, the way in which teixobactin kills germs is unique and shows great promise; recently, super-strong drug resistant bacteria have caused increased concern in the health community, with no clear solution in sight – this could be a valuable weapon. Also, the way in which they discovered the antibiotic shows great potential. The device has the potential to reveal further undiscovered antibiotics: it enables ‘unculturable’ microbes to thrive in the lab, and so makes it easier to discover bacteria that naturally produce compounds deadly to other pathogens.
“The technology is very cool,” says Gerard Wright, a biochemist at McMaster University in Hamilton, Canada, who was not involved with the study. “Nobody knew if these bacteria produced anything useful.
A Major Breakthrough

When you look at things in a broader context, the discovery method is even more exciting than the finding itself. A new antibiotic can only go so far, but a new way of finding antibiotics… now that’s a story! Most antibiotics we use today were discovered by scientists in the earlier part of the 20th century, and there’s been no new discovery for almost 3 decades. The antibiotics we know so far are:
1928 – Penicillin
1932 – Sulfonamides
1943 – Streptomycin
1946 – Chloramphenicol
1948 – Cephalosporins
1952 – Erythromycin and Isoniazid
1957 – Vancomycin
1961 – Trimethoprim
1976 – Carbapenems
1979 – Monobactams
1987 – Lipopepitides
As researchers combed through more and more samples, it became harder and harder to find new things; here’s how this team did it. First of all, they understood that the bacteria they were trying to grow simply don’t grow in a lab. Like most organisms, bacteria also take cues from their outside environment – if they don’t receive natural environmental cues, they simply don’t divide; so researchers had to take the bacteria in a different environment.
They suspended them in a lump of agarose (a sugar polymer) and then surrounded the polymer with a semi-permeable membrane which would not let cells through, but would let small molecules through. Then, they transplanted the entire system back into the soiland only then were they able to grow and mutate the bacteria in the lab.
“What most excites me is the tantalising prospect that this discovery is just the tip of the iceberg,” said Mark Woolhouse, professor of infectious disease epidemiology at the University of Edinburgh. “It may be that we will find more, perhaps many more, antibiotics using these latest techniques.”
It it also proves efficient in humans, then we’ll be likely seeing clinical trials in just a couple of years.
Journal Reference: Losee L. et al. A new antibiotic kills pathogens without detectable resistance. Nature (2015) doi:10.1038/nature14098