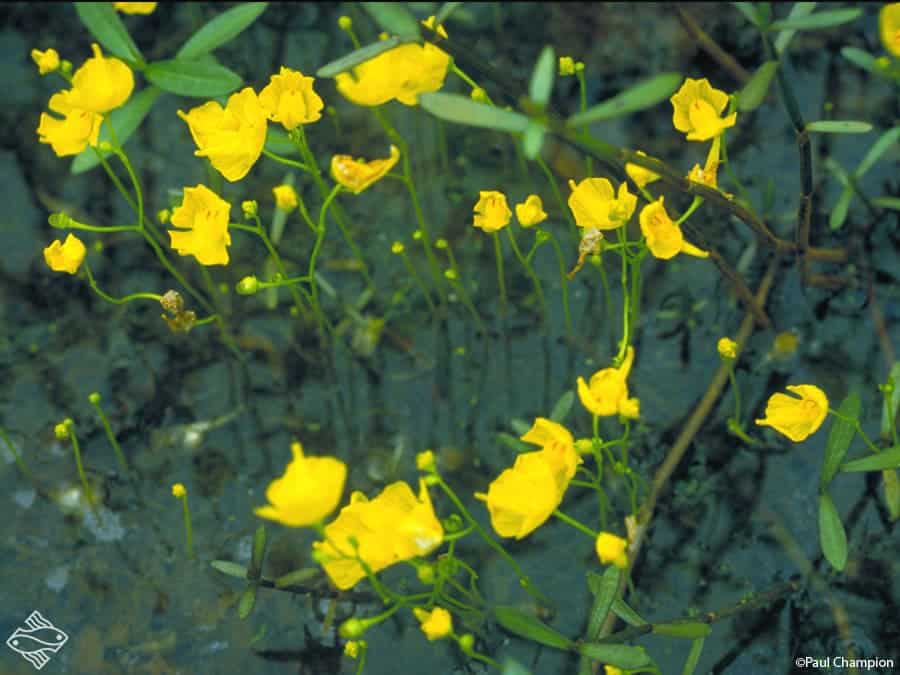
The carnivorous bladderwort (Utricularia gibba) is a carnivorous plant which occurs fresh water and wet soil. Recently, the plants took biologists by surprise by having a huge number of genes, despite a fairly small genome. The plant is six times smaller than the grape for example, but has 28,500 genes, compared to the 26,300 of the grape.
The genome is the genetic material of any living organism, encoded either in DNA or, for some viruses, in RNA. Typically, you’d expect a smaller genome to contain less genes, but it’s exactly the other way around for the bladderwort. This carnivorous plant somehow managed to cram a lot inside a very limited space – it’s a genetic architecture which has never before been seen.
“The story is that we can see that throughout its history, the bladderwort has habitually gained and shed oodles of DNA,” said Victor Albert, one of the researchers, in a news release. “With a shrunken genome, we might expect to see what I would call a minimal DNA complement: a plant that has relatively few genes-only the ones needed to make a simple plant. But that’s not what we see.”
Victor Albert of the University at Buffalo has been studying this issue for some years now. In 2013, he found that the bladderwort lacks “junk DNA” – portions of a genome sequence for which no discernible function has been identified. In other words, the plant somehow managed to shed all its useless DNA and only keep the valuable parts; only 3% of its DNA is “junk DNA”, compared to 90% for humans, for example (this doesn’t mean that 90% of our DNA is useless, but we haven’t found out its functions yet – still, there’s a good chance that a significant part will be confirmed to be junk DNA).
The plant is editing its own DNA at an incredibly fast pace – Albert and his colleagues believe that the plant’s genome has duplicated entirely at least three times. But instead of just adding all of those redundant genes to its DNA (the way crops like wheat have), it has kept cutting out the chaff.
“It turned out that those rates of evolutionary turnover — especially the rate of loss — was incredibly high compared to other plants,” Albert said. “The genome was subjected to some heavy duty deletion mechanisms.”
It seems logical that in time, when you have this extremely high pace of editing, the useless bits are quickly sorted out.
“When you have the kind of rampant DNA deletion that we see in the bladderwort, genes that are less important or redundant are easily lost,” said Albert. “The genes that remain-and their functions-are the ones that were able to withstand this deletion pressure, so the selective advantage of having these genes must be pretty high. Accordingly, we found a number of genetic enhancements, like the meat-dissolving enzymes, that make Utricularia distinct from other species.”
So the most important genes are kept, the useless ones are sorted out, and it’s not clear what happens to the marginally useful ones. Still, it’s extremely exciting to find this type of result – it basically shows that evolution can also take place (in one form or another) at a genetic level.
“It’s the kind of thing we were really hoping to see,” Albert said.
Still, it’s not clear exactly why this is happening – and why is it happening especially to this plant. In future research, Albert and his team hope to figure out the factors which led to this unexpected development – especially considering that other plants live in the exact same type of environment and compete for the same type of resources.
Journal Reference: Lorenzo Carretero-Paulet, Pablo Librado, Tien-Hao Chang, Enrique Ibarra-Laclette, Luis Herrera-Estrella, Julio Rozas and Victor A. Albert. High gene family turnover rates and gene space adaptation in the compact genome of the carnivorous plant Utricularia gibba. Mol Biol Evol (2015) doi: 10.1093/molbev/msv020.







