Biological systems have recently attracted the attention of mathematicians and computer scientists, who have been turning everything from quantum processes to RNA into logic gates. But before you get paranoid about DNA controlling Skynet, you have to know that these systems have only been successful on a small scale, calculating square roots on four-bit numbers, and each calculation took over five hours.
Today’s issue of Science jumps over small scale demonstrations, going directly into showing how a form of DNA computing can perform a calculation with up to 130 different types of DNA molecules involved. Despite its lack of speed, the system is incredibly flexible – flexible enough to allow the use of compilers and debugging circuitry.
Thing is, even though this technology isn’t exactly well suited for calculations, it can be integrated into biological systems, taking their input from a cell and feeding the output into biochemical processes. The authors of the papers are a biologist and a computer scientist from Caltech.
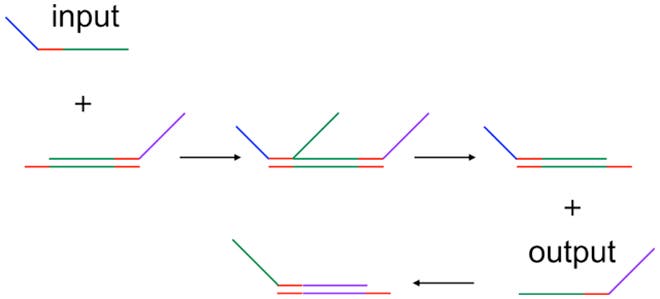
The system relies on what they have called “seesaw” logic gates, displayed above. The central feature of these gates is a stretch of DNA that can base-pair with many different molecules, allowing them to compete for binding. Even once a molecule is base-paired, it can be displaced; short “landing” sequences on either side allow a different molecule to attach, after which it can displace the resident one.
Because of the simplicity of logical operations and the fact that the rules of DNA base pairing are so straightforward they were able to generate a computerized model that told them what DNA molecules to acquire. In order to prove that it works, they constructed a system which calculated the floor of the square root of a four-bit binary number. The operation required 74 different single-stranded molecules of DNA, and during the time in which the operation was running, 130 different double stranded molecules existed in the same test tube.
A large number of biomolecules could be used as input here, including DNA, RNA, and small molecules. The authors also indicate that they have an idea on how to dramatically speed the process up – by using arge DNA scaffolds to assemble gates in close proximity to each other, ensuring that reactions take place quickly and require far less DNA to be used.
Via Wired, pictures via Ars Technica
Was this helpful?