With this technique, we can study cellular processes in unprecedented detail, researchers says.
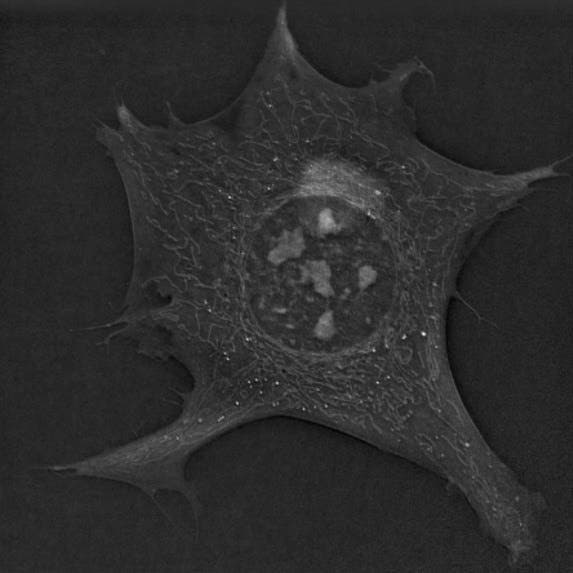
Studying cells is never easy. During the imaging process, cells can be damaged by light, while color dyes that make visualization easier can interfere biochemically. But in a recent study, researchers found a way to image cells in unprecedented detail without damaging them.
The technique combines image analysis with a 3D microscopy technique, and it produces more than just a pretty picture — it offers “access to an uncharted field of biological dynamics and describes a unique set of simple computer-vision strategies” researchers say.
“The ability to not only observe, but to measure dynamics of undisturbed living cells, through the application of quantification techniques to holo-tomographic microscopy, is likely to lead to the discovery of fundamentally new cell behaviors, and a new understanding of previously studied phenomena as well,” said lead author Mathieu Frechin of Swiss company Nanolive.
The technique relies on a new form of microscopy called holo-tomographic microscopy (HTM). HTM incorporates both holography and tomography. It splits a light beam into two, allowing only one part of it to pass through the sample. When the two parts are reunited, the differences in their waveform are used to infer various properties of the sample (based on its refractive index). Then, the process is repeated with the sample tilted at a slight angle, again and again, until the resulting images can be combined in a three-dimensional picture of a cell.
The method doesn’t involve any contrast dyes and it completely preserves the integrity of the cell, and it can be used to study a multitude of cellular processes, the researchers note.
In the study, researchers used this approach to highlight little-known processes about the cell. For instance, they showed that in the run-off to mitosis (one of the two types of cellular division, in which the nucleus divides in two), the entire nucleus rotates between 80 and 700 degrees over a period of a few minutes or a few hours. They also obtained quantitative data about the flux of fat within cells, measuring the formation and flow of fat droplets.
This type of approach can help us shed new light on a number of previously unknown processes, allowing researcherches to step into “uncharted field of biological dynamics,” the team says.
Journal Reference: Sandoz PA, Tremblay C, van der Goot FG, Frechin M (2019) Image-based analysis of living mammalian cells using label-free 3D refractive index maps reveals new organelle dynamics and dry mass flux. PLoS Biol 17(12): e3000553. https://doi.org/10.1371/journal.pbio.3000553






