A single virus making the jump from animals to humans can spark disaster, as we’ve seen so well over the past year. If viruses roam freely in animals, this raises the risk of the viruses mutating and ultimately making their way to humans. A new study found that this problem may be much more widespread than we thought — but there are ways we can prepare.
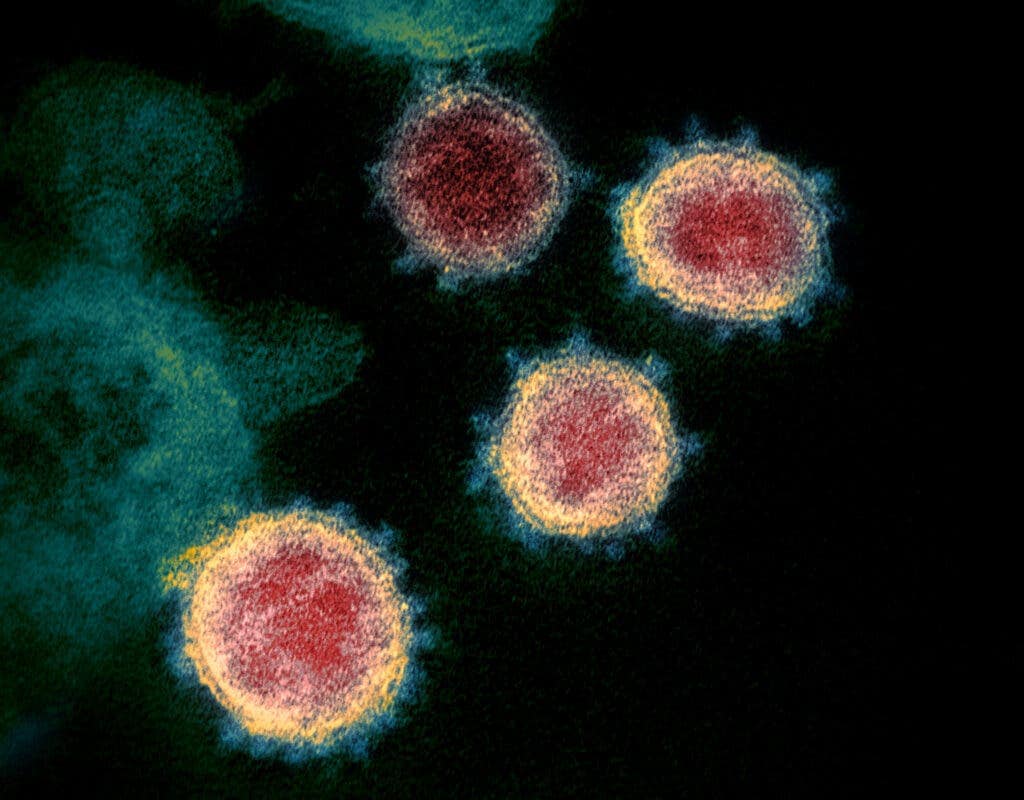
The COVID-19 pandemic started when a coronavirus made the jump from an animal (we’re still not 100% certain which one) to humans. At first glance it may seem like it came out of the blue, but the warning signs were there all along — not just with the SARS and MERS outbreaks, but also with studies that highlighted the risk of similar coronavirus outbreaks.
Now, researchers have highlighted another concerning trend, finding that there are at least 11 times more associations between mammalian species and coronavirus strains.
“This means that for every species that we have observed being infected with a specific coronavirus, there are another 11 different sets of wild mammal species and coronavirus infection that we haven’t yet observed. In short, there’s both a lot more and a lot more diverse coronavirus infections in wild mammals than we know about,” said Marcus Blagrove, one of the study authors, to ZME Science.
“We are not suggesting that recombination and production of new coronaviruses is something we should be worried about happening immediately,” Blagrove adds, emphasizing that this is a long-term trend. “This is a more long-term timescale.”
The study’s first author is Maya Wardeh, a specialist in biological data mining. Wardeh, Blagrove, and veterinary epidemiologist Matthew Baylis looked at the potential scale of novel coronavirus generation in both wild and domesticated animals. The goal was to identify likely mammalian hosts where the next coronaviruses may be generated over the next few years or decades, Blagrove explains in an email to ZME Science. If we know what the high-risk species are, we can keep an eye on them and identify potential pandemic viruses before they even jump to humans. This would serve as an effective warning system to either prevent future pandemics or at the very least, give us a headstart to work on effective vaccines and treatments early on.
New coronaviruses can emerge when different strains co-infect an animal, causing the viral genetic material to recombine, so infection patterns can help us understand when and where new coronavirus strains are likely to happen.
The research team looked at 411 strains of coronavirus and 876 potential mammalian host species using a machine-learning approach to predict relationships between the strains and species. They found that some species among the “usual suspects” — species previously associated with coronavirus outbreaks (such as horseshoe bats, palm civets, and pangolins), but also some new ones.
“Many previous and current studies are looking at which existing viruses can ‘jump’ from one species to another. Rather than looking at existing viruses, we look at where new coronaviruses may come from. In the last question, we talk about how new coronaviruses can be produced by viruses exchanging genetic material. We highlight that this can happen in many more mammalian species than we currently realize. Should human-infecting coronaviruses (such as SARS, and SARS-CoV-2) recombine with one of the many non-human coronaviruses, this can generate new viruses, with different diseases, that can affect humans in the future. This is how SARS (and probably SARS-CoV-2, it takes a long time to prove!) emerged.”
This means more species are sharing their viruses, which in turn means individual viruses can likely infect more mammals than expected. For humans, this is not a concern directly, but if these viruses start “meeting each other”, it can become a problem as they become more likely to jump.
“The problem with this is when two different strains of coronavirus meet each other in a host, then can produce new viral strains with attributes from each ‘parent virus’, in a similar way that two animals produce children with genetic material from each parent. This process is called homologous recombination.”
The overall scale of interactions suggests that viruses are coming into direct with each other much more than we realize, and it’s something we should be keeping an eye out for. The researchers predict 40 times more mammalian species can be individually infected with a wide range of coronaviruses than previously believed.
But the study doesn’t only raise an alarm: it also offers some practical solutions, such as keeping potential host species away from each other. Since some of these species are also domestic, protective measures also apply to agriculture, Blagrove concludes.
“Some aspects of animal welfare are particularly relevant to virus spread and, in this case, the generation of new strains. Our work highlights high-risk species of mammal which can be infected with large numbers of different coronaviruses. We would recommend minimizing the chances of these species coming into contact with more of these viruses, for example by minimizing the intensive farming of these animals and keeping them separate from other high-risk species so as to limit virus spread and thus chances for recombination. We also recommend surveillance of the particularly high-risk species to identify any new viruses early.”
The study has been published in Nature Communications.






