Researchers used new techniques to analyze the microbial diversity of ancient gut contents from two latrines: in Jerusalem, Israel, and Riga, Latvia. According to the findings, medieval gut bacteria was quite different to the one we have today.
Over the years, researchers have started to pay more and more attention to the human gut biodiversity. A healthy, resilient gut microbiome supports your immune system and may provide important health benefits, but we’re only now starting to look into the bowels of the problem.
A growing body of literature is associating changes in our microbiome to changes in our modern lifestyle, suggesting that conditions such as inflammatory bowel disease, allergies, and obesity may be intertwined with gut health. To find out more about how our gut microbiome has changed compared to pre-industrial times, a team of researchers analyzed a source of valuable information: latrines.
It’s an important source of information, but also a challenging one to analyze, says Piers Mitchell of Cambridge University, who specializes in the gut contents of past people through analysis of unusual substrates.
“Microscopic analysis can show the eggs of parasitic worms that lived in the intestines, but many microbes in the gut are simply too small to see,” comments Mitchell. “If we are to determine what constitutes a healthy microbiome for modern people, we should start looking at the microbiomes of our ancestors who lived before antibiotic use, fast food, and the other trappings of industrialization.”
It’s such a weird approach that Kirsten Bos, a specialist in ancient bacterial DNA at the Max Planck Institute for the Science of Human History, didn’t even think it was possible at first.
“At the outset we weren’t sure if molecular signatures of gut contents would survive in the latrines over hundreds of years. Many of our successes in ancient bacterial retrieval thus far have come from calcified tissues like bones and dental calculus, which offer very different preservation conditions. Nevertheless,” says Bos, “I was really hoping the data here would change my perspective.”
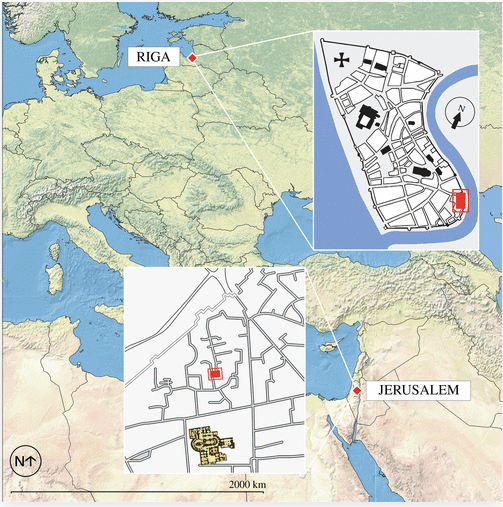
As the team got to analyzing sediment from the latrines in Jerusalem and Riga (from the 14-15th century), things got very interesting. The first challenge was distinguishing the bacteria that came from the gut contents of people from that naturally present in the environment.

Researchers were successful in this approach, identifying a wide range of microorganisms, including many known to inhabit the intestines of modern humans.
The two latrines shared common characteristics, but they exhibited noted differences compared to what you’d expect from a gut microbiome today. It’s not exactly clear what is causing this, but medieval microbiomes seem quite different from other ones.
Susanna Sabin, a co-author of the study, notes:
“We found that the microbiome at Jerusalem and Riga had some common characteristics — they did show similarity to modern hunter gatherer microbiomes and modern industrial microbiomes, but were different enough that they formed their own unique group. We don’t know of a modern source that harbors the microbial content we see here.”
It’s hard to draw any major conclusion from only two datapoints, but since the method seems viable, researchers are now considering using it on multiple archaeological sites to draw broader conclusions.
“These latrines gave us much more representative information about the wider pre-industrial population of these regions than an individual faecal sample would have,” explains Mitchell. “Combining evidence from light microscopy and ancient DNA analysis allows us to identify the amazing variety of organisms present in the intestines of our ancestors who lived centuries ago.”
The challenges still remain, however. Archaeologists need to first find suitable sites and then trial the method.
“We’ll need many more studies at other archaeological sites and time periods to fully understand how the microbiome changed in human groups over time,” says Bos. “However, we have taken a key step in showing that DNA recovery of ancient intestinal contents from past latrines can work.”
“It seems latrines are indeed valuable sources for both microscopic and molecular information,” concludes Bos.
Journal Reference: Susanna Sabin, Hui-Yuan Yeh, Aleks Pluskowski, Christa Clamer, Piers D. Mitchell, Kirsten I. Bos. Estimating molecular preservation of the intestinal microbiome via metagenomic analyses of latrine sediments from two medieval cities. Philosophical Transactions of the Royal Society B: Biological Sciences, 2020; 375 (1812): 20190576 DOI: 10.1098/rstb.2019.0576






