Just as whole organisms have an evolutionary history that can be traced backwards in time, revealing varying degrees of relatedness to other organisms through common ancestors, so too does each gene have an evolutionary history that can be traced back through time. Many modern genes are part of gene families, each gene in the family having similar, yet distinct functions, and originally derived from a common gene ancestor. A new study presented in the January 12, 2016 issue of the Proceedings of the National Academy of Science (PNAS), uncovers one method evolution takes to go from a single primitive gene to multiple genes, each with unique abilities and discrete functions.
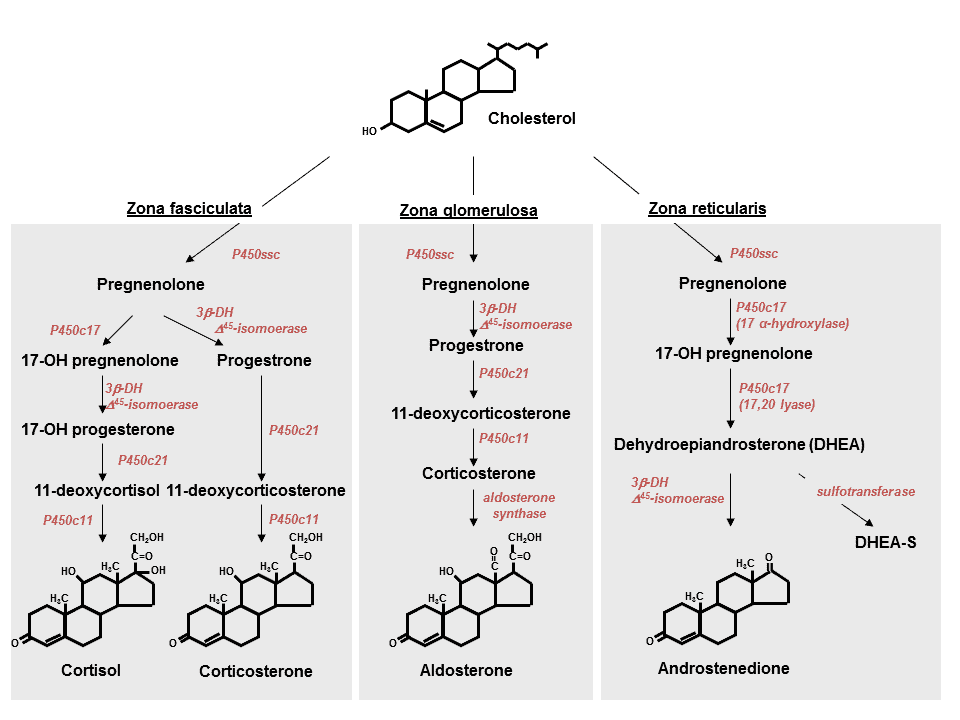
The study looked at the evolution of the Steroid Hormone Receptor family of genes. Steroids are essential molecules in animals, that are chemically derived from cholesterol, to produce a large variety of hormones, from cortisol and aldosterone (products of the adrenal glands), to sex hormones like estrogen and testosterone. When one of the steroid hormones enters the cell it exerts its influence on DNA through a Steroid Hormone Receptor. It is the Steroid Hormone Receptor that binds to the appropriate place in the DNA, to either activate or repress other genes. For example, cortisol can have an anti-inflammatory effect by binding to the Glucocorticoid Receptor, suppressing genes in cells of the immune system, essentially turning these cells off. It is this property of corticosteroids that make them so useful for treating pro-inflammatory conditions, like those seen during acute injury or chronic autoimmune disease. On the other hand, aldosterone can have a pro-inflammatory effect, by binding to the Mineralocorticoid Receptor, and activating a different set of genes.
Both the Glucocorticoid Receptor and the Mineralocorticoid Receptor are members of the Steroid Hormone Receptor gene family. There are six known members in all, but only the Glucocorticoid Receptor can repress gene activation. The other members of the Steroid Hormone Receptors are gene activators. The study focused on how the Glucocorticoid Receptor developed the capacity to inhibit gene expression.
The team found that all members of the Steroid Hormone Receptor gene family appear to be derived from a distant ancestral gene whose function was promiscuous, in the sense that this Steroid Hormone Receptor had properties that could both up regulate and down regulate genes. Which ever action predominated may have been influenced by other factors in the cellular environment. The team reconstructed the DNA sequences of the ancestral Steroid Hormone Receptors at several points along evolutionary history. What they found was that gene duplication events lead to multiple copies of the ancestral gene, which allowed the new separate copies to evolve down different pathways, acquiring distinct functions from the common ancestor gene. The common ancestor of all Steroid Hormone Receptors showed both gene up regulating action, along with a mild degree of gene down regulating activity, when tested in cell culture.
After the gene duplication event, one copy of the gene became the common ancestor to the Glucocorticoid and Mineral Corticoid Receptors, while the other copy became the common ancestor of the Progesterone and Androgen Receptors, which began specializing their function as they gained new mutations. Analysis revealed that going from the common ancestor of all Steroid Hormone Receptors to the common ancestor of the Glucocorticoid and Mineral Corticoid Receptors, happened with the acquisition of two simple base changing mutations. Those two mutations were neutral, meaning that they did not change either the up regulating or down regulating ability of the receptor – it had basically the same activity as the common ancestor of all Steroid Hormones – but it did produce a slight change in the 3-D shape of the molecule.
A later mutation in a single base lead to complete loss of the Mineralocorticoid Receptors ability for gene down regulation. Interestingly, it was the single base mutation, in combination with the previous two neutral mutations that were necessary to lose the down regulating action. This seems to suggest that, so called neutral mutations, may still be important to derive new traits, if they prime the system for further change.
The Glucocorticoid Receptor, on the other hand, gained several more mutations that further enhanced its gene down regulating ability, above and beyond the ancestral gene. It was the occurrence of several gene duplication events, this last one being in the common ancestor of all bony vertebrates, that provided the raw material to evolve separate genes with more discriminating functions, losing the promiscuity of a single “jack of all trades” gene to gain a family of genes that became specialists, allowing a more finely tuned and regulated system. By performing X-ray crystallography and Nuclear Magnetic Resonance analysis of the various receptors molecules, the team further showed that changes in up or down regulating activity coincided with minor differences in the 3-D structure of the receptors, affecting their interaction with the DNA.
Journal Reference:
Distal substitutions drive divergent DNA specificity among paralogous transcription factors through subdivision of conformational space. Proc Natl Acad Sci U S A. 2016 Jan 12;113(2):326-31. doi:10.1073/pnas.1518960113. Epub 2015 Dec 29. Hudson WH, Kossmann BR, de Vera IM, Chuo SW, Weikum ER, Eick GN, Thornton JW, Ivanov IN, Kojetin DJ, Ortlund EA.
Was this helpful?



